We are the group led by Prof. Qu. Here shows our research topics.
Bacteria swimming/swarming
Individual bacterium swims in planktonic enviroment follow a “run-and-tumble” mode, while bacterial collective motion (swarming) does not.
- How this transition happens? What is the underlying physics?
- Can we harvest energy from such a swarming system, which exhibits chaotic motion? If so, how?
- Can we create smart, soft, biological robots from real biological systems?
Microscopic view of bacteria swarming
Macroscopic view of bacteria swarming in the self-developed imaging platform
Algae Swimming
In addition to studying bacteria, we also investigate the locomotion of algae, specifically Chlamydomonas reinhardtii. Our research includes exploring phenomena such as algae accumulation near solid-liquid interfaces and their oriented motion along patterned surfaces. These observations raise several key questions:
- What mechanisms drive this accumulation phenomenon? What are the underlying physical principles?
- Are these effects purely the result of hydrodynamic interactions?
- How can we develop practical applications to eliminate algae accumulation when it has negative impacts?
Long time tracking of Chlamydomonas reinhardtii realized by 3D real-time tracking microscope
Chlamydomonas reinhardtii swimming near sinusoidal wave like surfaces
Active matter system
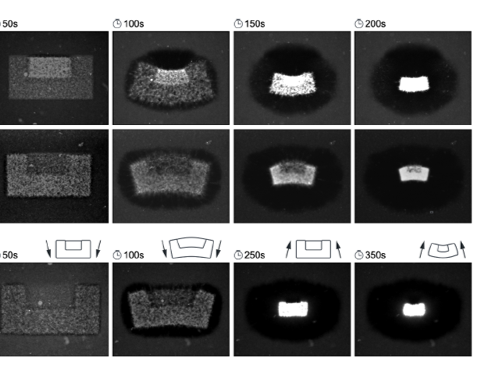
Our active matter system consists of purified microtubules and light-activatable motor proteins that crosslink and organize the microtubules into distinct structures upon illumination. By shining light on them, these motor proteins are activated and they will connect the scattered microtubules together, causing the system to contract. This can help us to achieve the functions of locomotion and reconfiguration which is related to some living systems. For our own system, we are interested in the shrinkage behavior, fluid flows caused by the shrinking.
We can achieve the bending behavior in the shrinking process by shining different intensities of light in different areas. The higher intensity can get higher density of microtubules. In the third case, the pattern of illumination varies in time to change the bending direction as the network contracts. The areas of the higher intensity and lower intensity switch.

The active matter system can also be used to capture and move cells (ongoing). The asteroid object in the video is the activated matter system. It can grab some cells it touches.
Apart from active matter systems including the purified microtubules and light-activatable motor proteins, we also investigate the bacteria avtive matter system applying methods in turbulence analysis.
Macroscopic model of bacterial flagella interaction and propulsion generation
Limited by micron-sized experimental objects, systematic manipulation of bacteria is complex and often impossible. Therefore, our group proposes a macroscopic experimental facility that can measure propulsion force and observe the flagella interaction based on previous research.
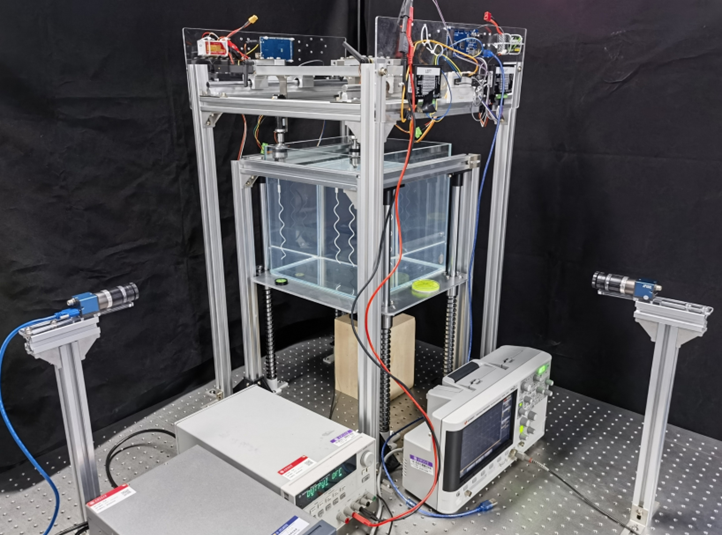

The macroscopic experiment system is designed to achieve four functions: propulsion force measurement, two-camera-synchronization, 2 DOF motor platform, and macroscopic helix models with bionic hooks. Based on this experimental system, typical “bundle” phenomenon of helices are observed. Also, the propulsion force generated by helices will be directly recorded via sensors with high precision.
Testing Results
3 Helices “Bundle” phenomenon
2 Helices “Bundle” phenomenon
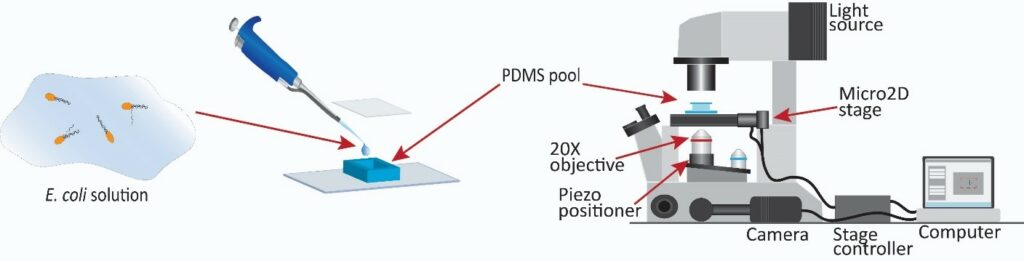
Real-Time 3D Digital Tracking Microscopy
To observe the swimming behavior of the cells, our group designed a 3D digital tracking microscope. It allows us to study the motility of individuals for a relatively long period (15 s on average) compared with more common stationary microscope stages.
The cells are observed using a Nikon Eclipse Ti2 Inverted Microscope with a CFI Plan Fluor 20XMI objective and PCO edge 5.5 sCMOS camera. A 2D translational stage (Prior) is used for tracking the cells in the 2D plane parallel to the glass slide. A piezo objective positioner (Physik Instrumente) is used to rapidly control the position of the focal plane in real time so that the cell always remains in the field of view. Images are acquired at 60 fps with a resolution of 640 pixels × 480 pixels. A real-time algorithm, written in C++ and OpenCV, detects the position (centroid) of a single cell in the image and moves the stage and objective to maintain the cell in focus and within the field of view.